dna
Brain cells snap strands of DNA in many more places and cell types than researchers previously thought.
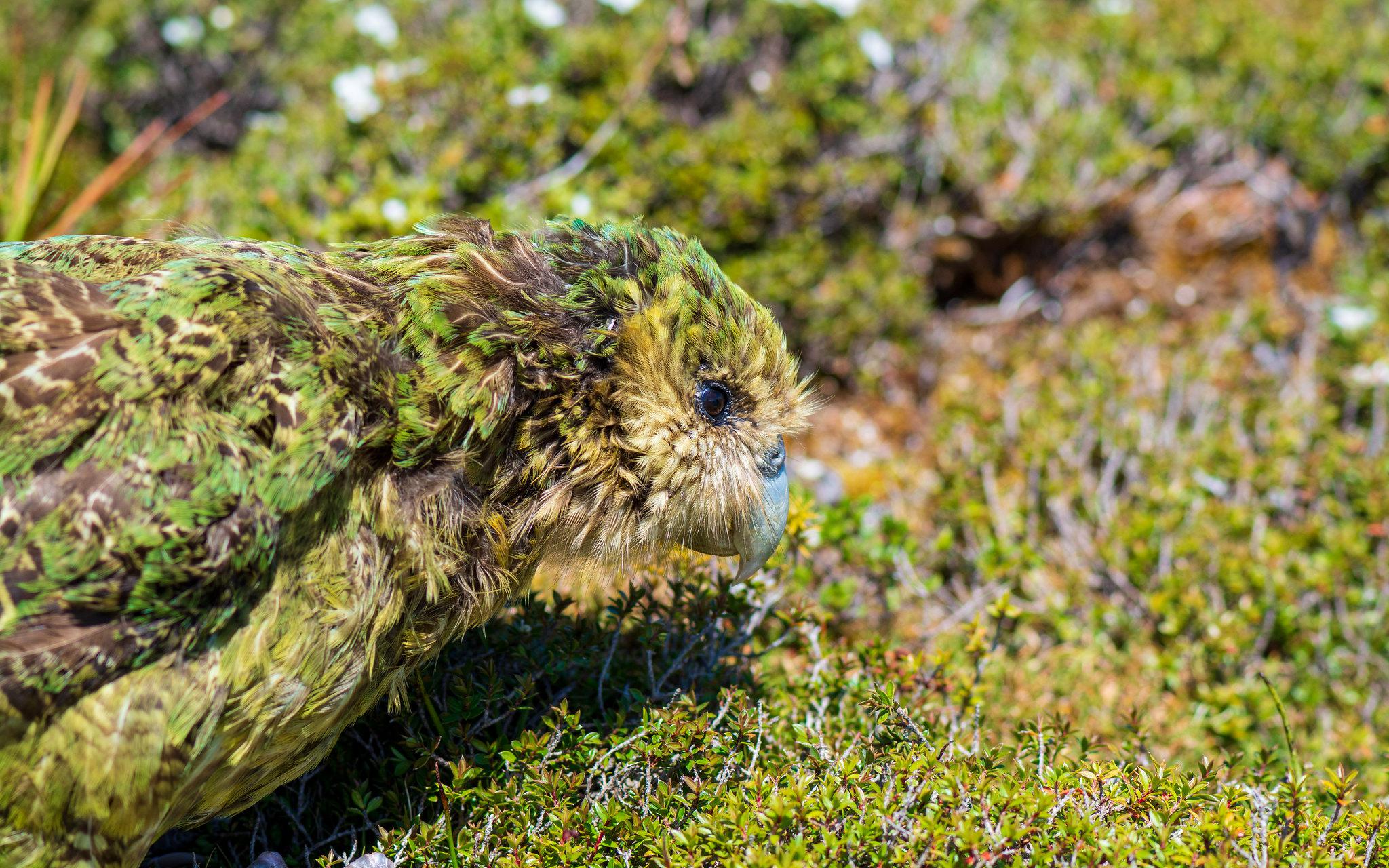
The Vertebrate Genomes Project may spell good news for the kakapo and the vaquita.
“The question is which are okay, which are not okay.”
▸
14 min
—
with
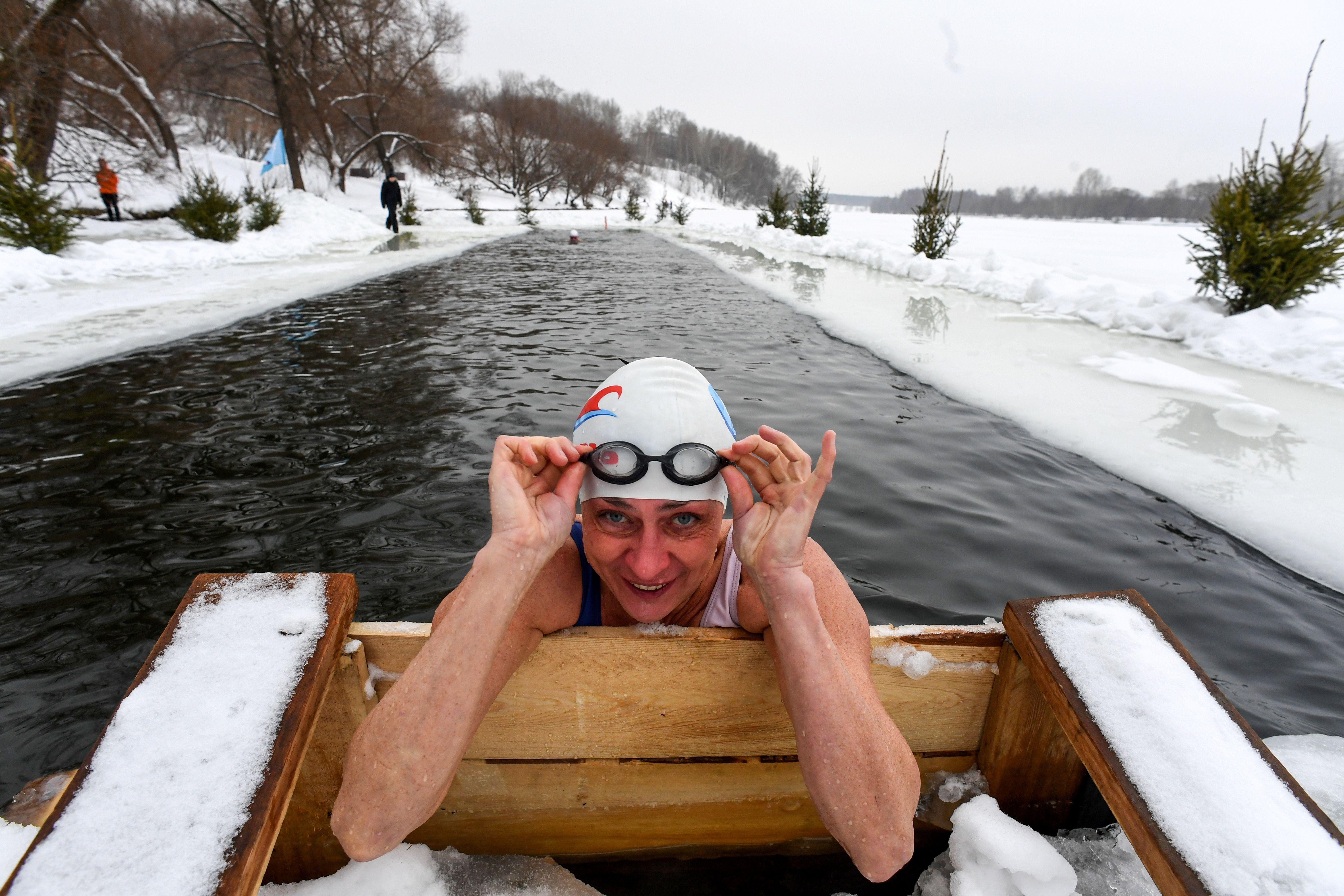
What makes some people more likely to shiver than others?
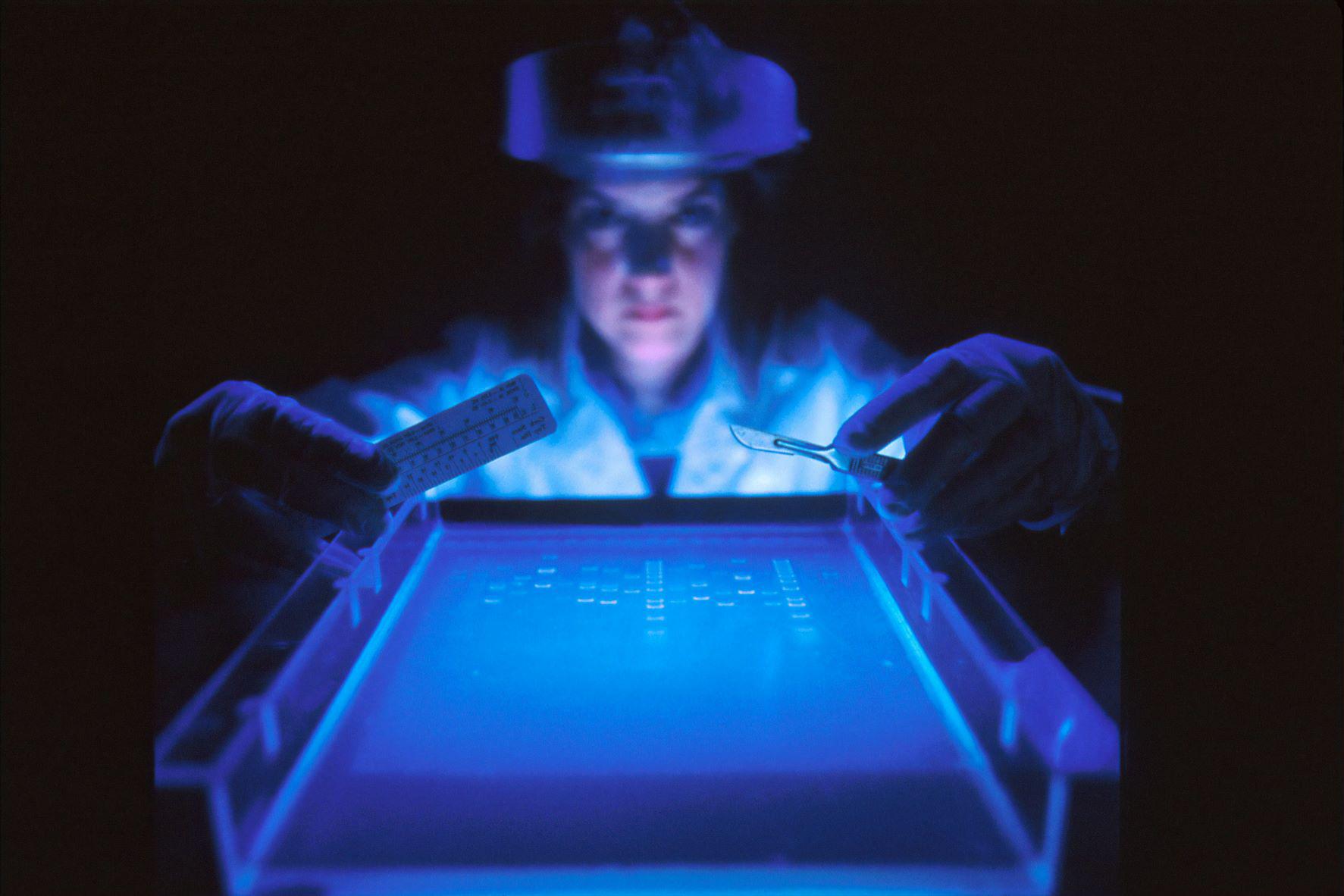
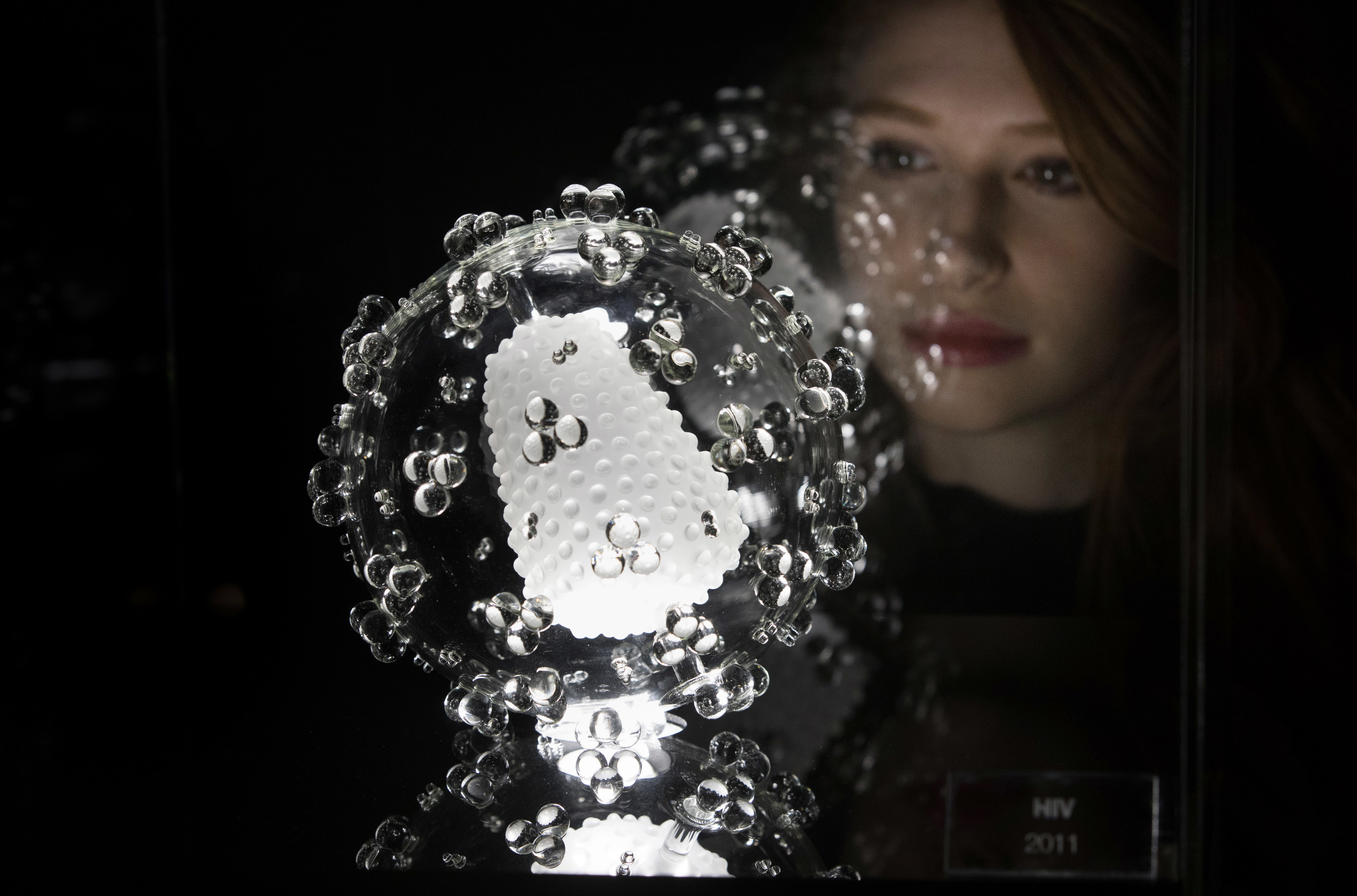
Scientists use high resolution microscopy and computer simulations to create first ever video of DNA movements.
The study found that people who spoke the same language tended to be more closely related despite living far apart.
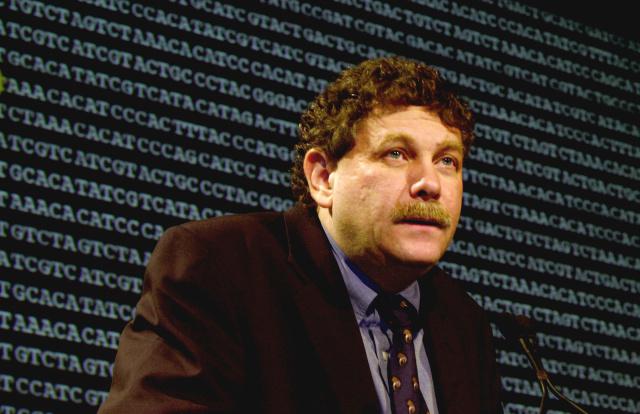
Dr. Eric Lander is a pioneer in genomics. What role will he play in the new administration?
Scientists find that an RNA-DNA mix may have created the first life on our planet.
Two new studies shed light on who first inhabited the islands, who replaced them, and how few people lived there.
There is a neurological link between serotonin levels and the brain’s ability to control impulses and patience levels.
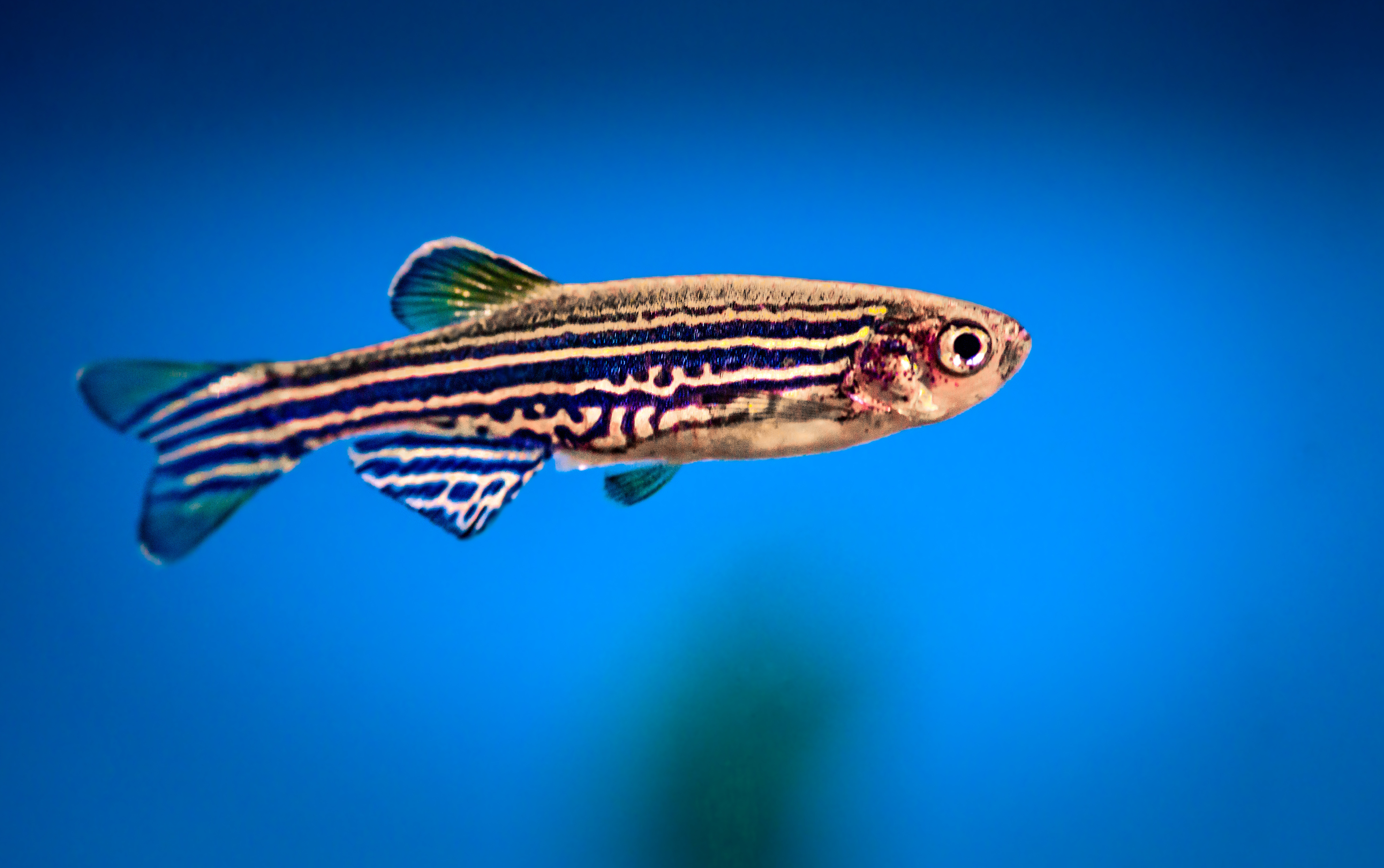
These tiny fish are helping scientists understand how the human brain processes sound.
The compound found in “magic mushrooms” has significant and fast-acting impact on the brains of rats.
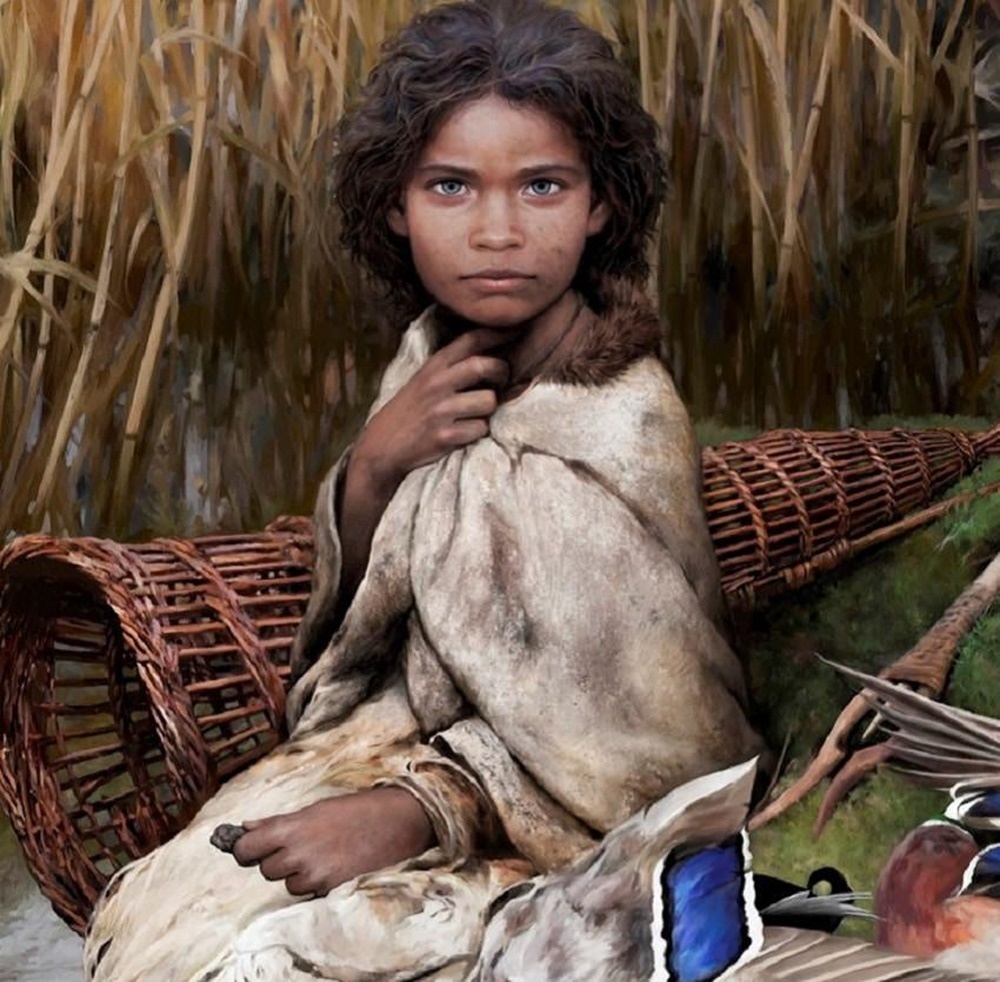
“The results change the perception of who a Viking actually was,” said project leader Professor Eske Willerslev.
DNA molecules are highly programmable.
Just how close are we to setting up camp on another planet? It’s complicated.
▸
17 min
—
with
Our family tree is complicated, and some of the branches are still unlabeled.
Is CRISPR the solution?
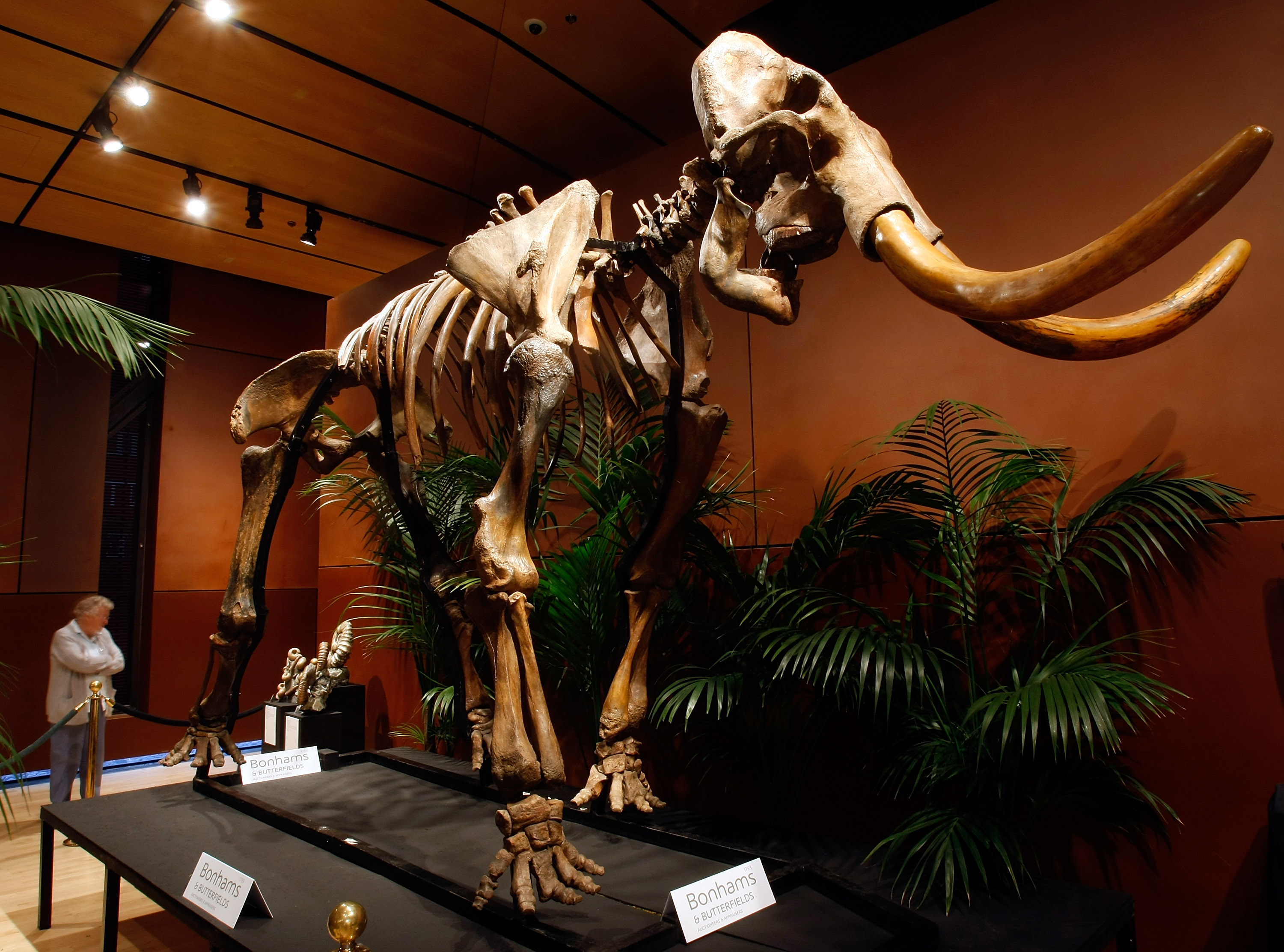
Scientists uncovered the secrets of what drove some of the world’s last remaining woolly mammoths to extinction.
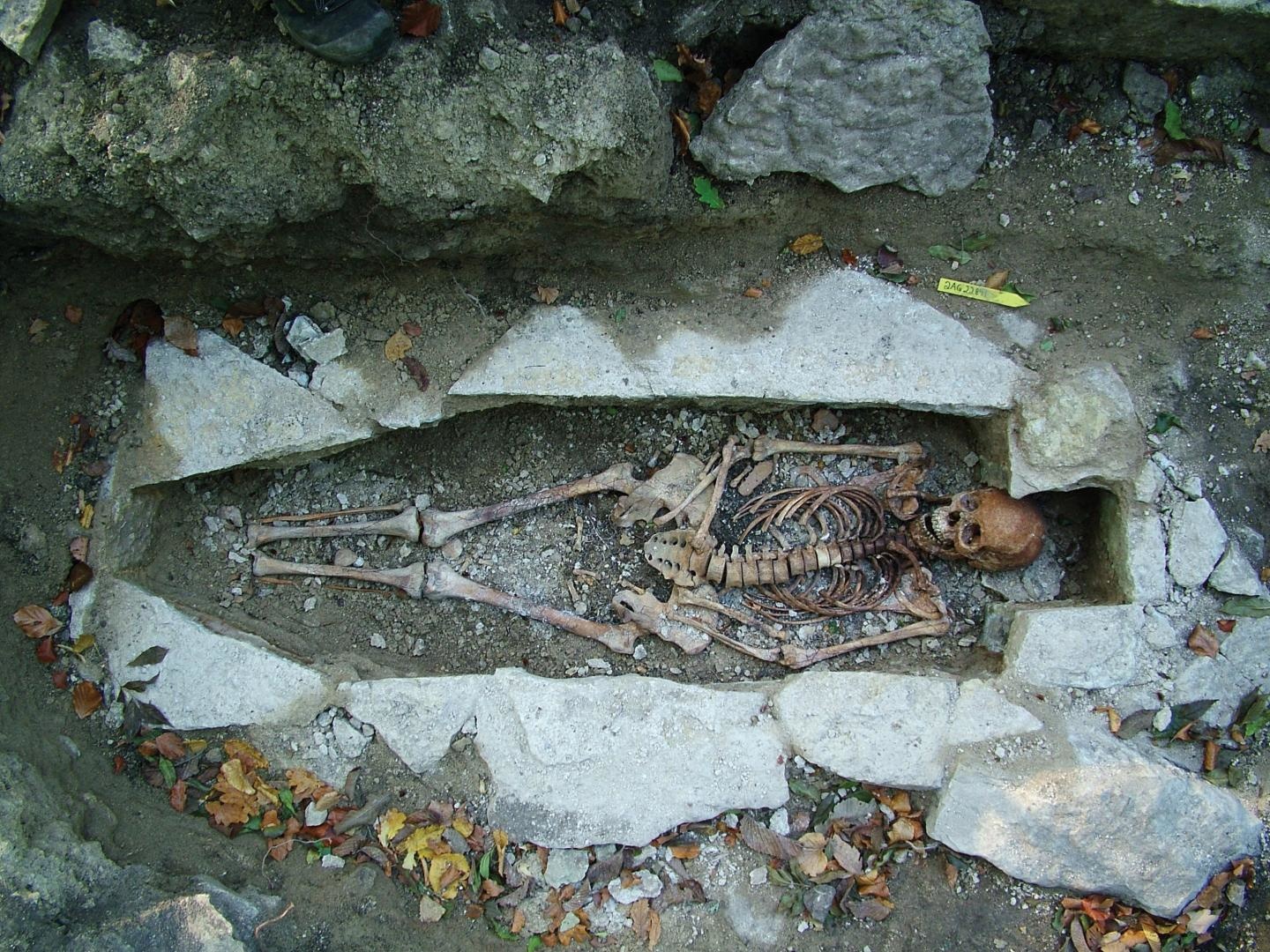
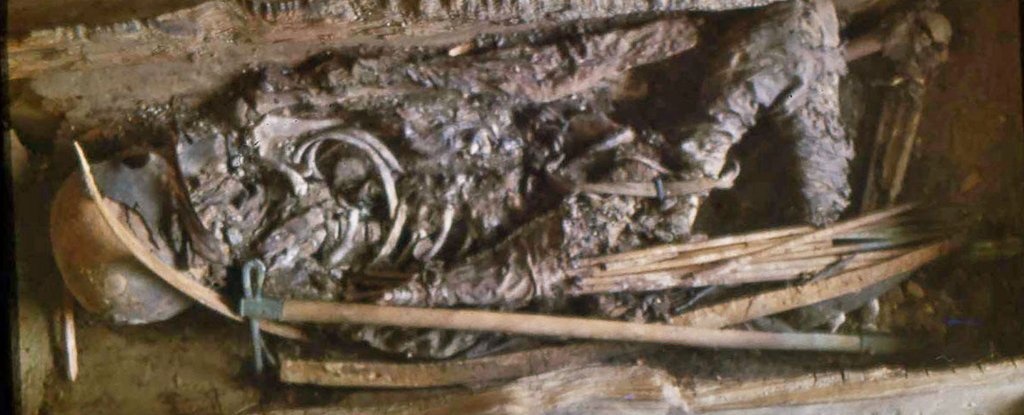
Thanks to modern technology, we can reexamine our assumptions about ancient warriors.
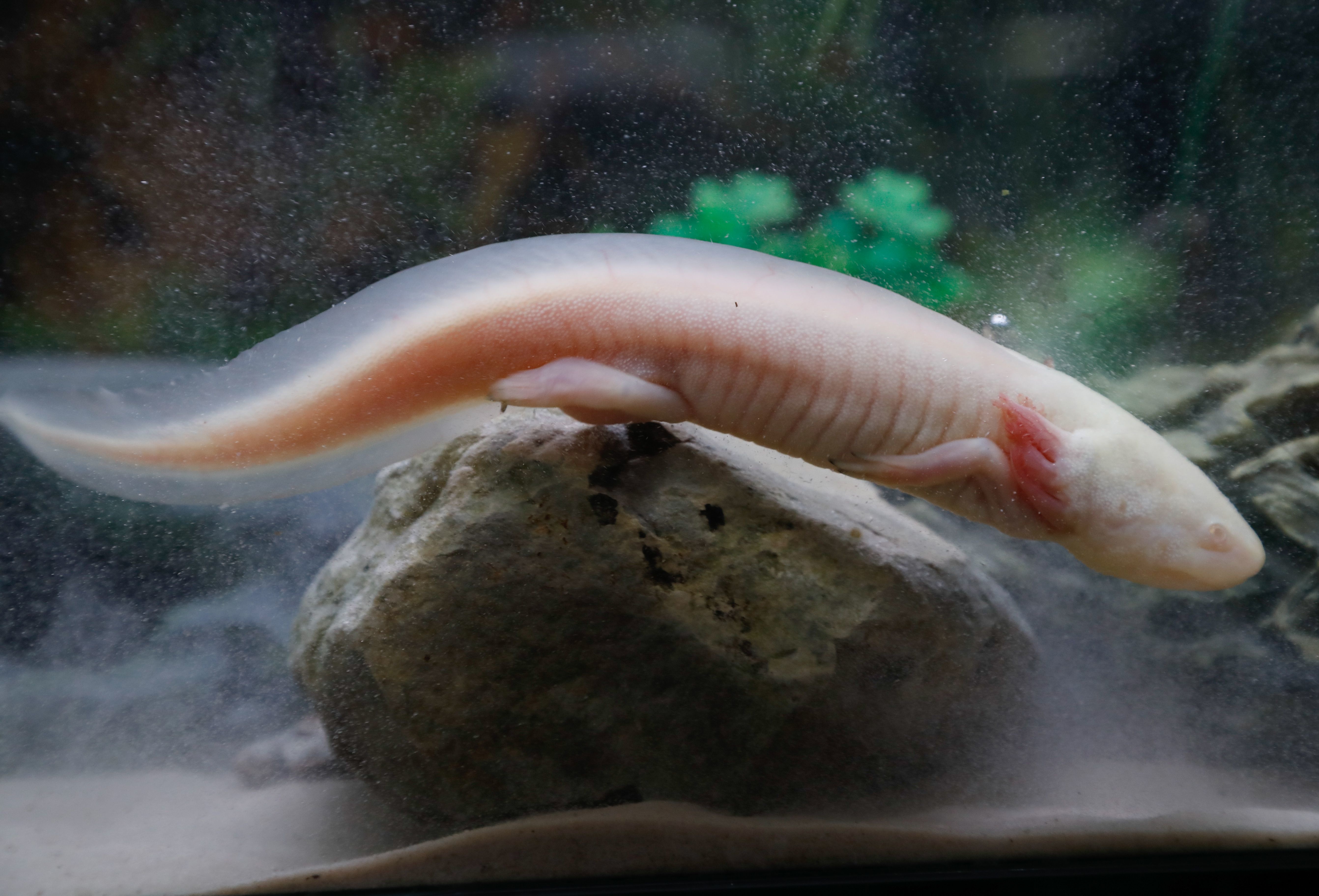
Researchers have recently discovered two of the genes that govern this weird-looking salamander’s ability to regenerate limbs, eyes, and even its brain.
If your New Year’s resolution was to get in shape, signing up for the marathon is a bad way to go about it.
All this from a wad of gum?
Learn where your ancestors are from, the breed of your rescue dog, and if your home is safe with these easy at-home kits.
A recent computer analysis found that millions of possible chemical compounds could be used to store genetic information. This begs the question — why DNA?
New forensic evidence and three other theories on London’s most notorious serial killer
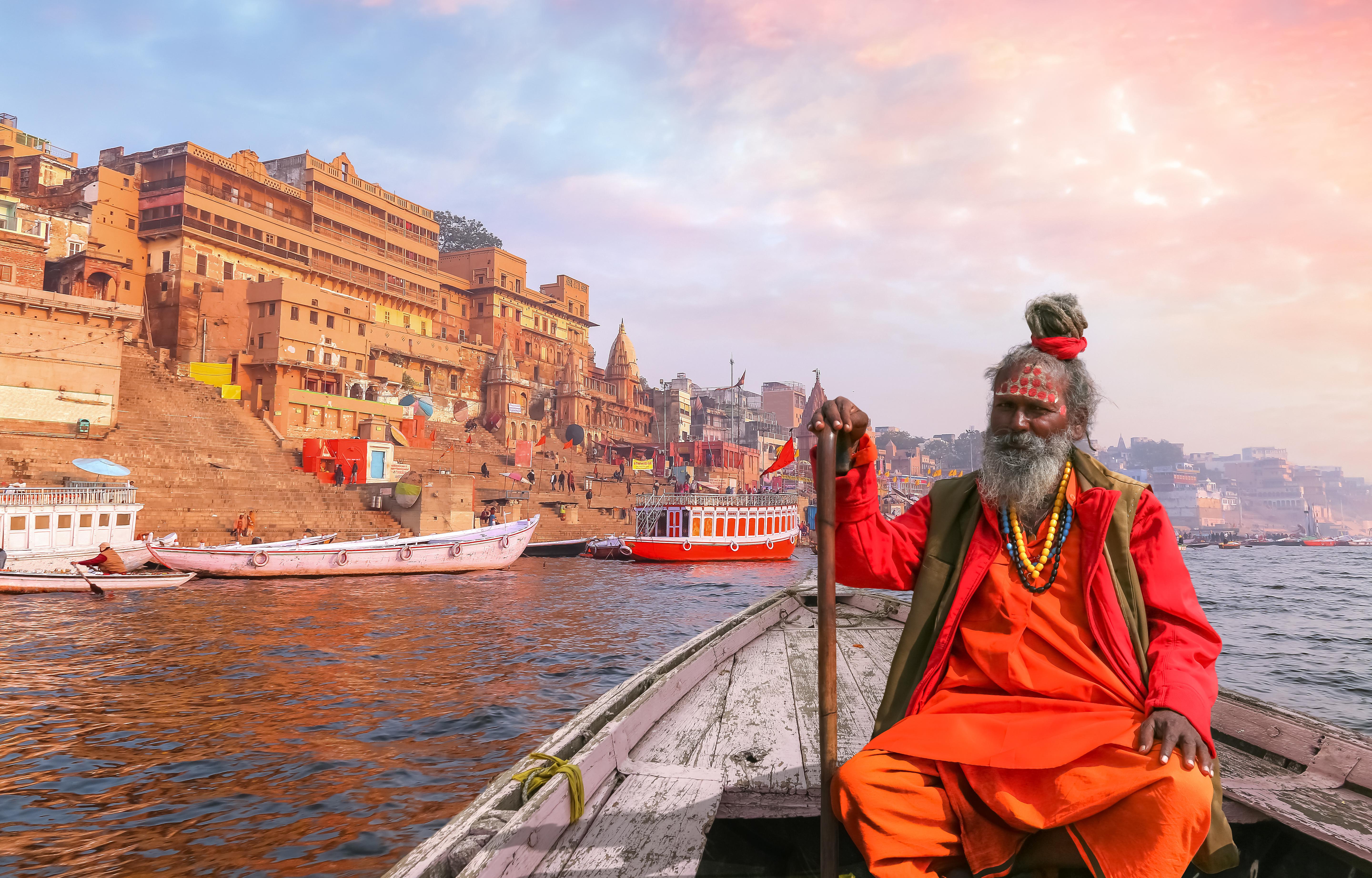
A DNA study looks for the home of the earliest modern humans.
A single typo in the “dark matter” of the genome drives multiple types of cancer.
While the blockbuster franchise might have given us a distorted view of science’s capabilities to address species extinction, new research might come close to “resurrecting” lost species’ DNA.
Some scientists think there may be a hidden, second form of life living right under our noses.
The question is no longer “can we” but “should we” edit human embryos.