genes
Porcine gene edits may allow such transplants without rejection.
How important is it to consider a romantic partner’s genetic profile before getting married?
The relationship between our genotypes and our psychological traits, while substantial, is highly indirect and emergent.
A new study shows how interbreeding of modern humans and Neanderthals boosted our genomes.
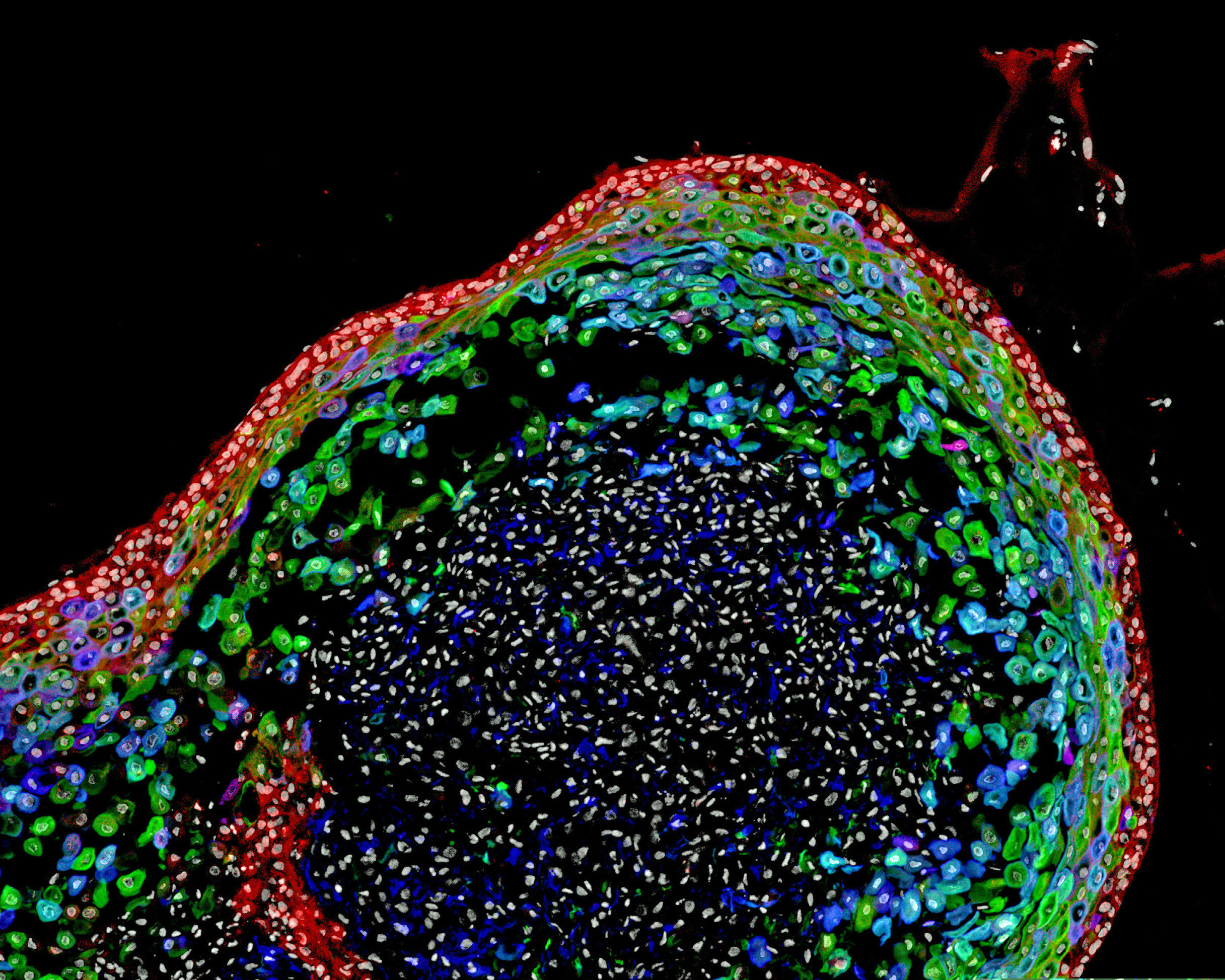
Scientists have grown a model human esophagus using pluripotent stem cells for the first time.
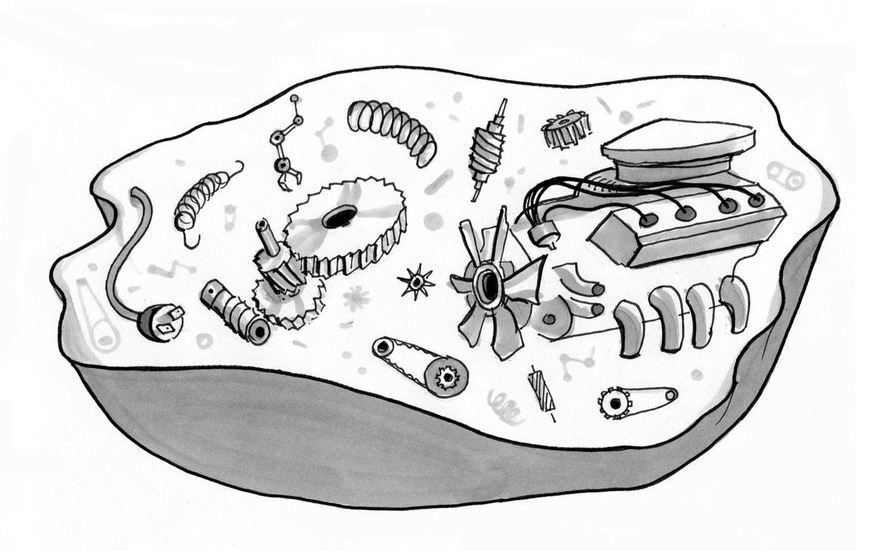
We can “read” genes with ease now, but still can’t say what most of them “mean.” To show why we need clearer “causology” and fitter metaphors, let’s scrutinize cars and their parts like we do bodies and genes.
First, let me tell you how smart I am. So smart. My fifth-grade teacher said I was gifted in mathematics and, looking back, I have to admit that she was […]
“Scientists should think like poets,” says E.O. Wilson, because new metaphors mobilize new thinking.
Funnily enough, some humans carry one of the very same genes.
A chorus of new science is showing that evolution has orchestrated life to leave no room for solos. A grander view of life is revealing higher-level, need-centric relational logic patterns (as in David Haskell’s The Songs of Trees).
Human minds are all powered by the same organ, so why do we have such strong preferences and diverse favorite things? Bill Nye lets us in on an example from his life.
▸
3 min
—
with
We are what we are because of genes; we are who we are because of memes. Philosopher Daniel Dennett muses on an idea put forward by Richard Dawkins in 1976.
▸
7 min
—
with
A new genetic study sees us getting gently stupider over time.
Scarcely noticed in the Eden story, there lurk fruitful scientific ideas about why biology generated morality.
This particular type of gene therapy gave mice younger bodies and 30% longer lifespans.
Why are we the way that we are – is it nature or nurture? This week, Bill Nye answers a question from Evan, who is having a science argument with his mom.
▸
2 min
—
with
Does life work like our technology? Is life under the hood just like a car sporting souped-up complexity?
Our picture of life is going through a major shift. Ed Yong’s book I Contain Multitudes reveals that a genome generally doesn’t contain all the genes an organism needs. Symbiosis isn’t rare, it’s the rule. And we’re just the icing on life’s vast microbial cake.