Scientists discover new DNA structure that’s not a double helix
In 1953, American biologist James Watson and English physicist Francis Crick published a scientific paper that revealed a groundbreaking discovery: the shape of DNA.
“This structure has novel features which are of considerable biological interest,” they wrote.
That turned out to be an understatement. The discovery of the double helix would go on to revolutionize the scientific world, leading to major developments in disease screening and treatment, food production, and forensic science.
Now, scientists have identified another structure of DNA.
In a paper published in Nature Chemistry, researchers from Australia describe the first-ever sighting of a DNA component—called the intercalated motif (i-motif)—within living human cells. The shape of the structure has been likened to a “twisted knot.”

“The i-motif is a four-stranded ‘knot’ of DNA,” said genomicist Marcel Dinger, who co-led the research. “In the knot structure, C [cytosine] letters on the same strand of DNA bind to each other—so this is very different from a double helix, where ‘letters’ on opposite strands recognize each other, and where Cs bind to Gs [guanines].”
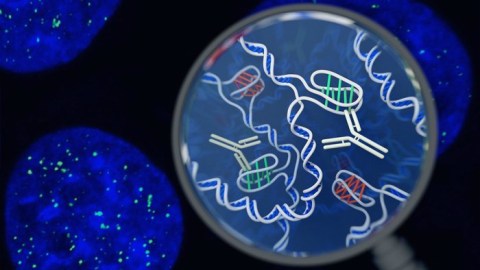
To identify the i-motif, which had been previously identified in vitro but never in living cells, the researchers developed antibody fragments dubbed “iMabs” that could recognize and bind to i-motifs in cells. The researchers added fluorescent dyes to the iMabs to make them easy to spot.
“What excited us most is that we could see the green spots—the i-motifs—appearing and disappearing over time, so we know that they are forming, dissolving and forming again,” said Dr. Mahdi Zeraati, the first author of the new paper.
The transient nature of i-motifs, along with the fact that they’re observed near areas of cells that control whether genes get activated, provides clues about their functions.
“We think the coming and going of the i-motifs is a clue to what they do,” said Zeraati. “It seems likely that they are there to help switch genes on or off, and to affect whether a gene is actively read or not.”
Dinger expressed a similar idea to the Australian Financial Review. “It may function as a volume dial in the genome that regulates activity of genes and can impact on the amount of a gene that is expressed in a cell.”
Zeraati said it is important for future research to uncover more about the functions of the i-motif and other DNA structures.
“These alternative DNA conformations might be important for proteins in the cell to recognize their cognate DNA sequence and exert their regulatory functions,” he told Science Alert. “Therefore, the formation of these structures might be of utmost importance for the cell to function normally. And, any aberration in these structures might have pathological consequences.”