genome
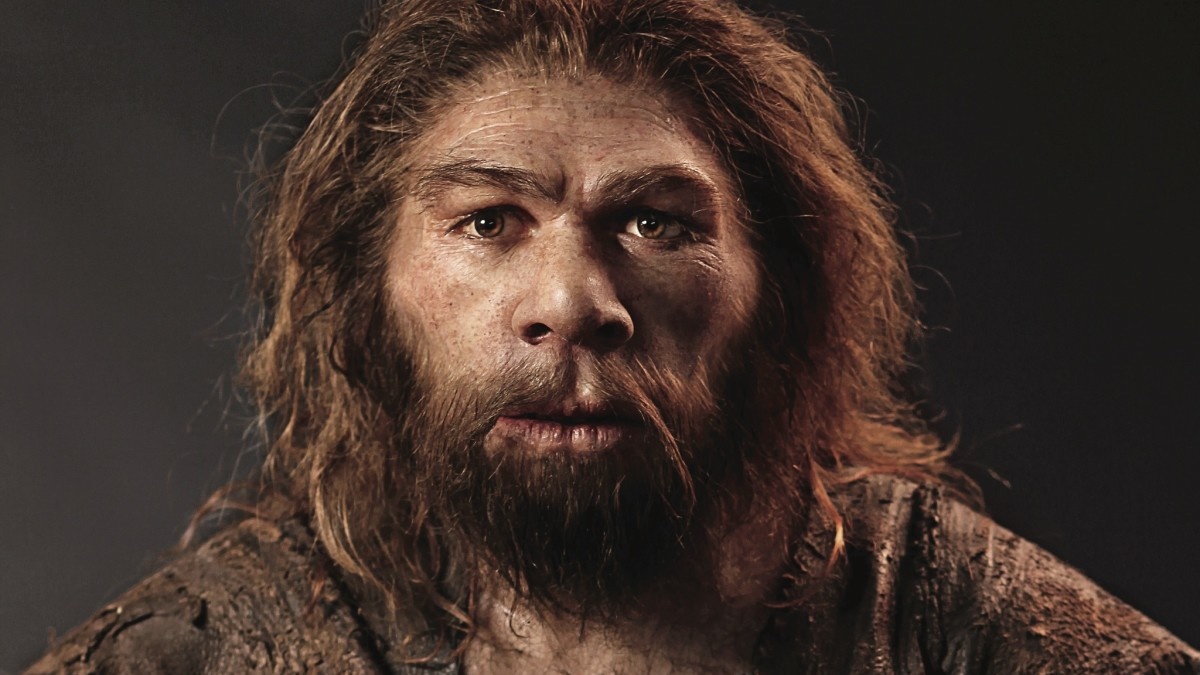
A new study shows how interbreeding of modern humans and Neanderthals boosted our genomes.
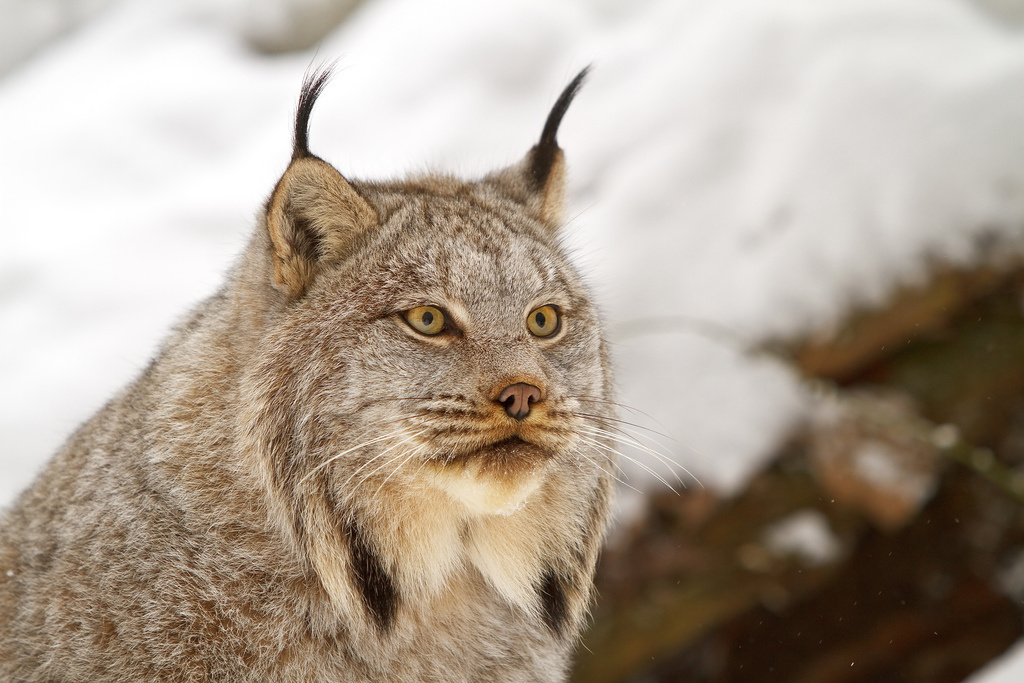
If you want to know what makes a Canadian lynx a Canadian lynx a team of DNA sequencers has figured that out.
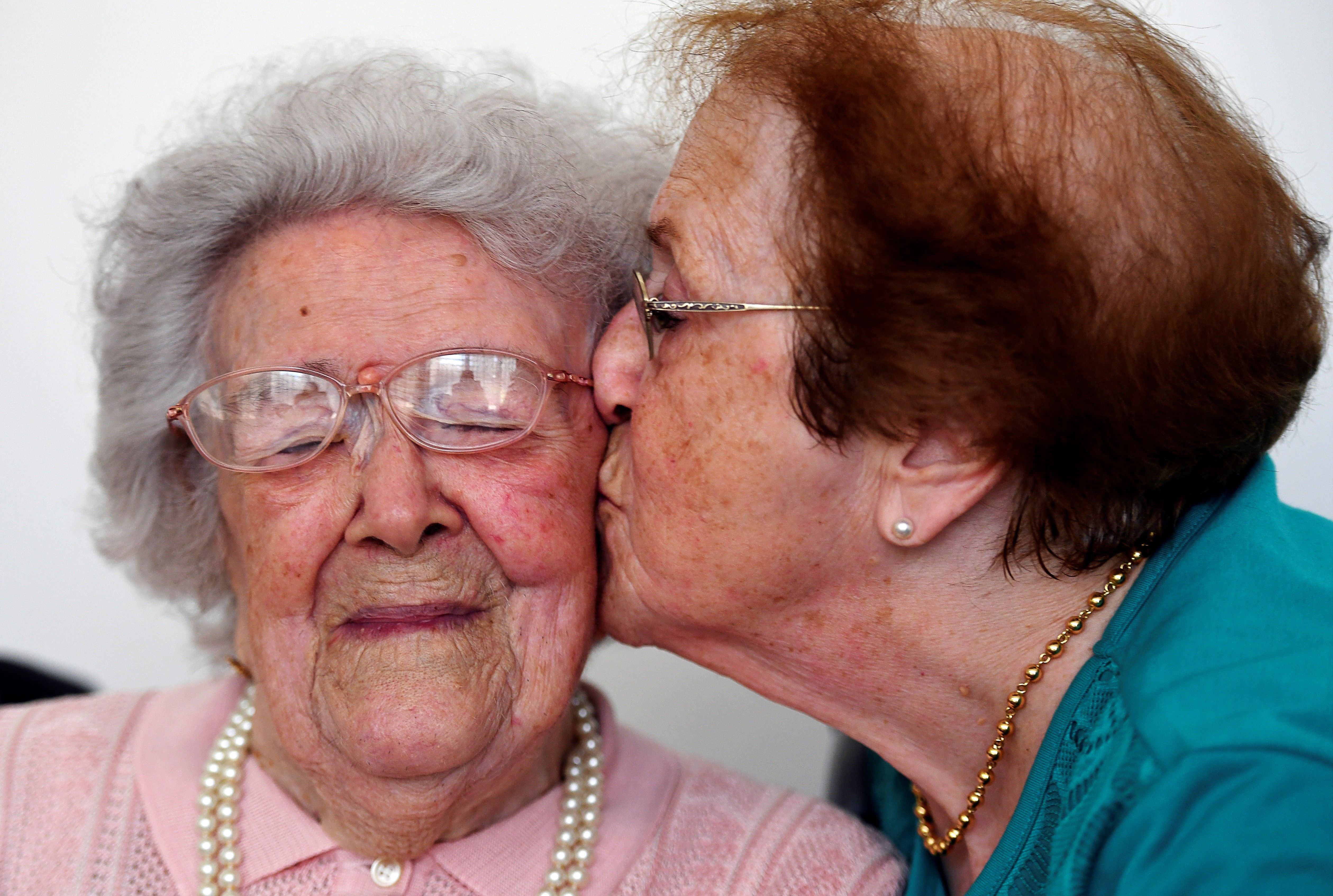
Scientists can now virtually reconstruction certain long-dead individuals, without the need for DNA samples from physical remains.
Find the right genes and we’ll have a way to prolong life and good health, perhaps indefinitely.
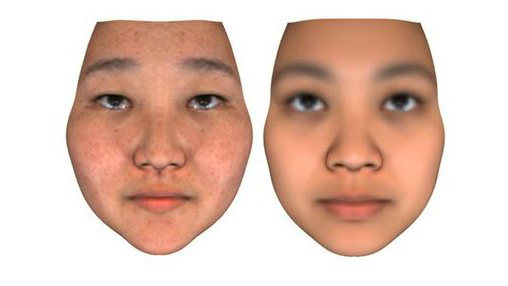
Researchers at Human Longevity have developed technology that can generate images of individuals face using only their genetic information. But not all are convinced.
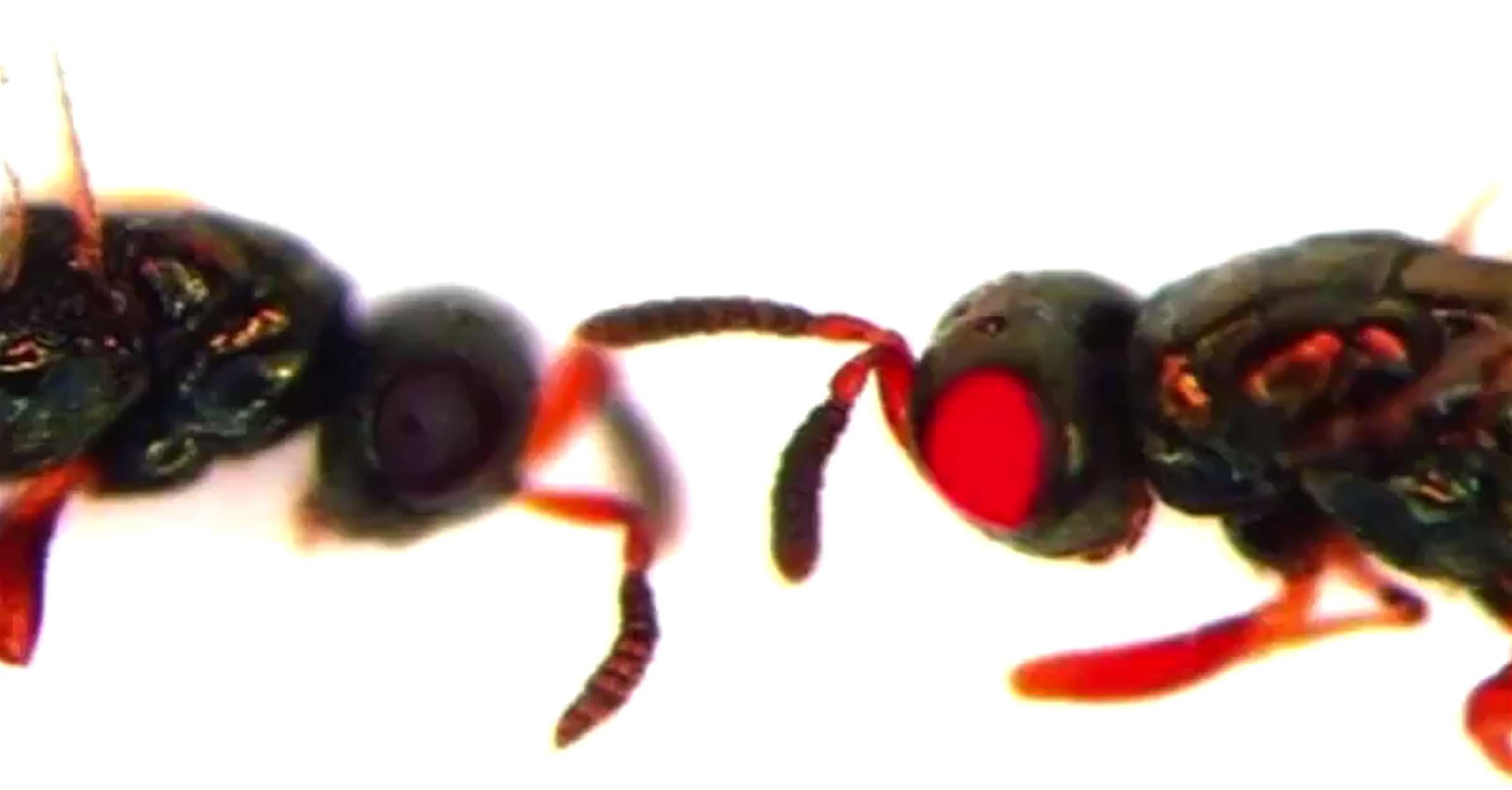
Researchers succeed in deleting key genes from ants, significantly modifying their behavior.
Being able to rewrite DNA as we wish could give us almost god-like power over all life on earth.