genomics
Technological advances in molecular biology could help fix the planet.
▸
2 min
—
with
Granted, genetic manipulation has been a dream for decades. Here’s what is different now.
Despite doubts from the healthcare industry, Watson recently identified a surprisingly high number of potential cancer treatments for real-life patients.
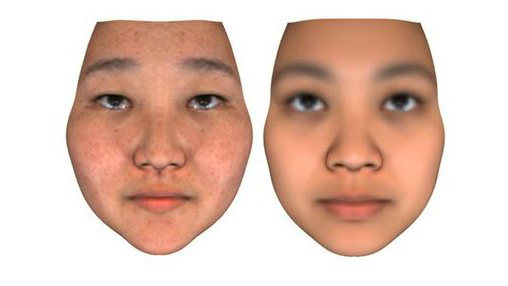
Researchers at Human Longevity have developed technology that can generate images of individuals face using only their genetic information. But not all are convinced.
No pep talks here, just a prediction by innovation expert Alec Ross that gene code and precision medicine is set revolutionize life the same way that computer code has.
▸
4 min
—
with