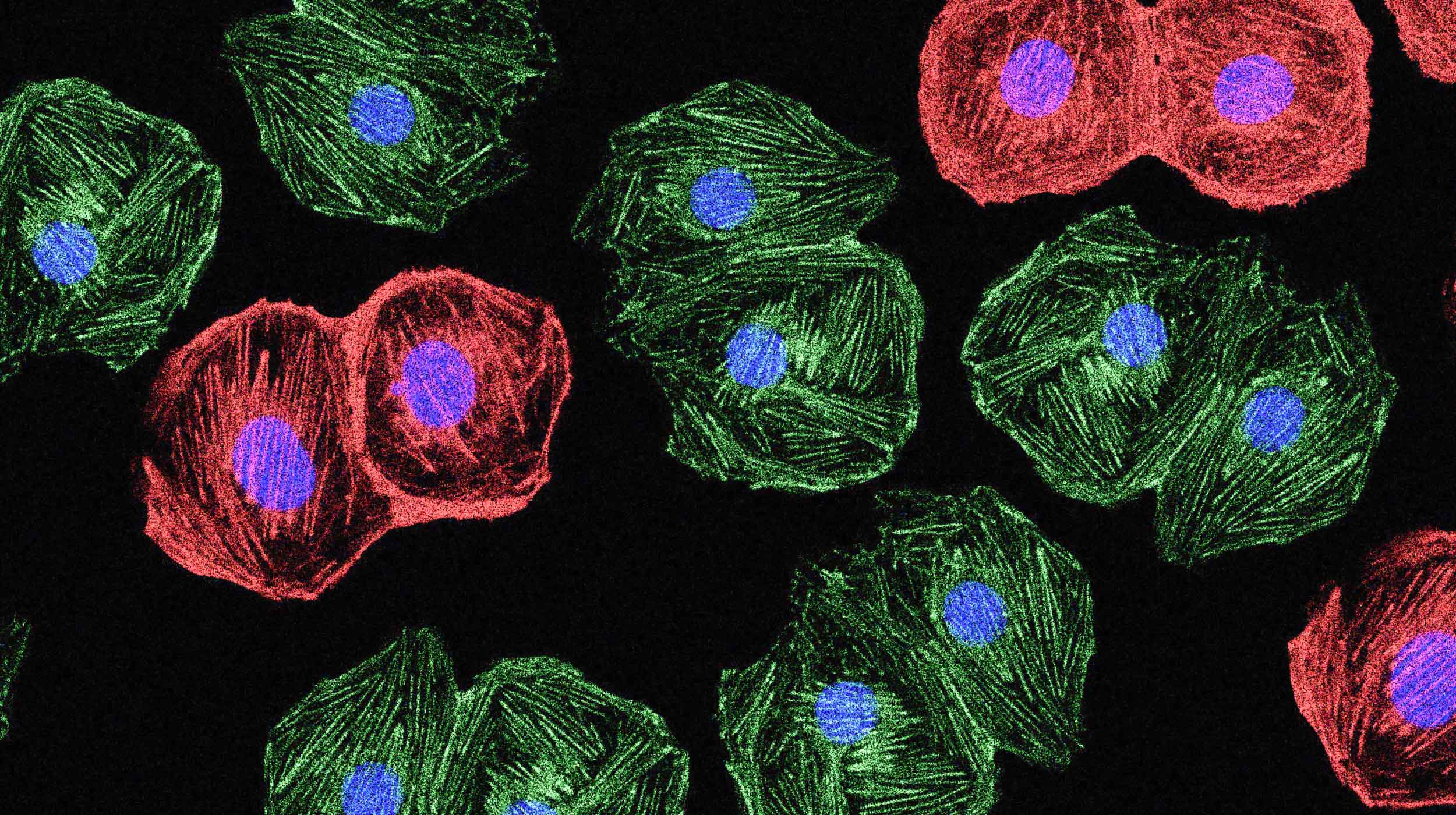
In the murky darkness, blue and green blobs are dancing. Sometimes they keep decorous distances from each other, but other times they go cheek to cheek — and when that happens, other colors flare.
The video, reported last year, is fuzzy and a few seconds long, but it wowed the scientists who saw it. For the first time, they were witnessing details of an early step — long unseen, just cleverly inferred — in a central event in biology: the act of turning on a gene. Those blue and green blobs were two key bits of DNA called an enhancer and a promoter (labeled to fluoresce). When they touched, a gene powered up, as revealed by bursts of red.
The event is all-important. All the cells in our body contain by and large the same set of around 20,000 distinct genes, encoded in several billion building blocks (nucleotides) that string together in long strands of DNA. By awakening subsets of genes in different combinations and at different times, cells take on specialized identities and build startlingly different tissues: heart, kidney, bone, brain. Yet until recently, researchers had no way of directly seeing just what happens during gene activation.
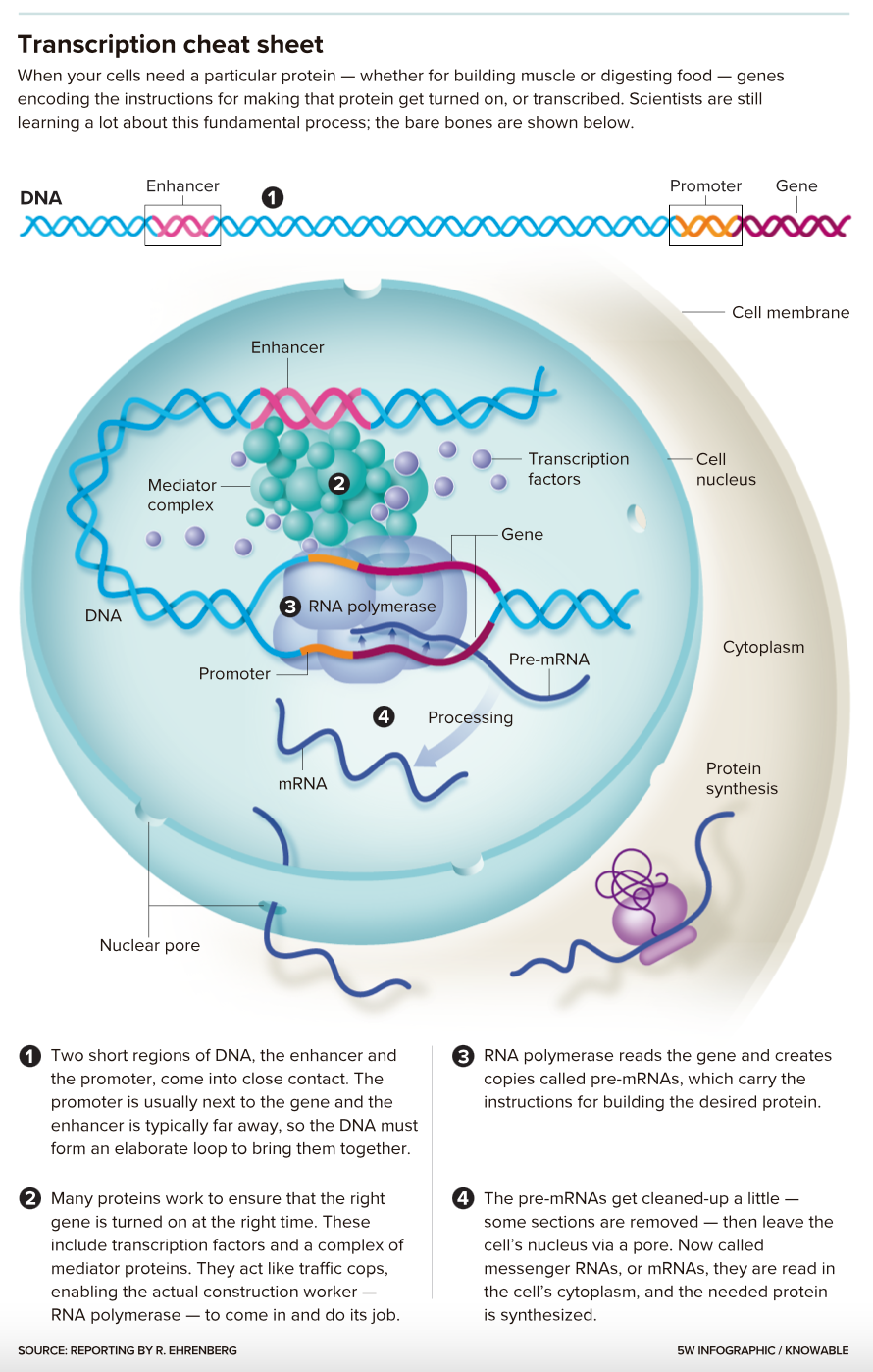
They’ve long known the broad outlines of the process, called transcription. Proteins aptly called transcription factors bind to a place in the gene — a promoter — as well as to a more distant DNA spot, an enhancer. Those two bindings allow an enzyme called RNA polymerase to glom onto the gene and make a copy of it.
That copy is processed a bit and then makes its way to the cytoplasm as messenger RNA (mRNA). There, the cellular machinery uses the mRNA instructions to create proteins with specific jobs: catalyzing metabolic reactions, say, or sensing chemical signals from outside the cell.
This textbook take is true as far as it goes, but it raises many questions: What tells a given gene to turn on or off? How do transcription factors find the right sites to bind to? How does a gene know how much mRNA to make? How do enhancers influence gene activity when they can be a million DNA building blocks away from the gene itself?
or decades, scientists had only blunt and indirect tools to probe these questions. Ideas about DNA, RNA and proteins came from grinding up cells and separating components. Then, in the 1980s, scientists began using a game-changing technique called FISH (short for fluorescence in situ hybridization) to see DNA and RNA directly, right in the cell. Other methods followed — microscopes with better resolution, new ways to tag (and thus track) players in this molecular symphony as it played out. Researchers could parse transcription as it happened, in detail.
Before, it was like trying to hear the symphony by looking at a static picture of the orchestra, says Zhe Liu, a molecular biologist at the Howard Hughes Medical Institute’s Janelia Research Campus in Virginia. “You would never figure out what they are playing,” he says. “You could never appreciate how beautiful the symphony is.”
Here’s a taste of what molecular biologists are learning by spying on this key, nanoscopic process — increasingly in real time, in living cells.
The life and times of transcription factors
Though scientists have long known that transcription factors dictate whether or not a gene powers up, it’s been mysterious how these proteins navigate the ridiculously crowded space in the nucleus to find their binding sites.
Consider that, uncoiled, the DNA in a human cell would run a meter or two long. The nucleus is about 5 to 10 micrometers in diameter, so the packaging of our genome is akin to stuffing a string that could wrap 10 times around the Earth inside a chicken egg, Liu says.
Researchers are just starting to tackle how this coiling and looping affects gene transcription. For one thing, they suspect it could help explain how enhancers can influence a gene’s activity from a great distance — because something far away when DNA is stretched out may be a lot closer when the genetic material is bundled up.
And if it seems miraculous that transcription factors know where they are going — well, most of them don’t. By tracking these proteins in a single cell over time, researchers find that they spend fully 97 percent of their life jiggling hither and thither, bouncing off of whatever bits of DNA they encounter until they luck out. (A few types may act as leaders, scanning the genome, latching on to their target and setting up the right conditions for a larger pack to follow.)
One would imagine, at least, that when a transcription factor finally found its binding site, it could stay stuck and do its job for hours. Scientists used to believe so from experiments with dead, preserved cells.
But studies on live cells show that’s far from true. Liu’s lab and others have shown over the past five years that transcription factors bind only for seconds, and that high concentrations of them congregate near the binding site, helping each other glom on. “It’s mind-boggling how transcription factors actually work,” Liu says.
And there are a lot of them: Up to 10 percent of the genes in a mammal carry instructions for making ones of different flavors. Recent evidence suggests that this affords huge precision to the cell. For any given gene, varied combinations of transcription factors can ramp up or tamp down the process, potentially making the system exquisitely tunable.
Hooking up at the polymerase party
If transcription factors are the gas pedal and brakes, the engine is RNA polymerase. In the basic model, RNA polymerase pulls apart a gene’s two strands, then slithers down one of them to make an mRNA copy of it. Turns out things are a hair more complicated.
Studies in mashed-up and preserved cells had hinted that many polymerase molecules cluster together to make this mRNA magic happen. But no one had ever seen such a clump in living cells, so no one knew how or when — or even if — the clumps formed. By attaching a fluorescing chemical tag to RNA polymerase in live cells, researchers saw multiple polymerases repeatedly group together for about five seconds — then scatter.
Last year, the same team of scientists spotted gatherings of other proteins as they congregated to help RNA polymerase do its job. These beasts — known as mediator proteins — form giant clusters numbering in the hundreds that join the RNA polymerases on the DNA.
The two gaggles seem to concentrate into distinct droplets, like blobs of oil in water. Then they fuse, perhaps creating a kind of self-assembling, cordoned-off transcription mill. A lesson from this? “Beyond the biochemistry, there are all these physical phenomena that may have a role in telling us how genes get turned on,” says biophysicist Ibrahim Cissé of MIT, who led the work.
Messenger RNA is made in fits and starts
For decades, researchers assumed that when a gene is active, transcription simply goes into “on” mode and cranks out mRNA at a steady clip. But a breakthrough technique called MS2 tagging, first developed in 1998 and still widely used, has radically changed that view.
Invented by cell biologist and microscopist Robert Singer and colleagues at the Albert Einstein College of Medicine in New York, MS2 tagging allowed scientists to see mRNAs in living cells for the very first time. (Key ingredients of the method come from a virus called MS2 — hence the technology’s name.)
In a nutshell, scientists use engineering tricks so that mRNA made from a specific gene bears distinctive structures called stem-loops. Through a second trick, those stem-loop locations are made to glow fluorescently so researchers can “see” mRNA from the gene of their choice whenever it is made and wherever it travels to, under a microscope and in real time.
Singer, who coauthored a 2018 article about mRNA imaging in the Annual Review of Biophysics, used MS2 tagging to show, with his colleagues, that the production rate of mRNAs from a gene fluctuates wildly over 25 minutes or so. It turned out that the size of these bursts doesn’t vary much, but their frequency does, and that’s what dictates how energetically a gene pumps out its mRNA product. Increasing or decreasing the rate of this transcriptional “bursting” may allow the system to ramp up or slow down a gene’s activity to meet the cell’s needs.
Researchers think that the on-off kinetics of transcription factors, meaning the rate at which they pop on and off of their binding sites, somehow regulates transcriptional bursting. But they don’t yet know how.
Trekking towards translation
Making mRNA is just the first step in a gene’s strutting its stuff. Next comes translating instructions in that mRNA to make proteins. For that to happen, the mRNA must journey out of the nucleus and into the cytoplasm where the protein-making factories reside.
Scientists had assumed that the cell’s molecular machinery carefully transported mRNA to the nucleus’s membrane and then pumped it out into the cytoplasm. Using the same MS2 method, Singer’s lab found that wasn’t so. Instead, mRNAs bounce around — “buzzing around in the nucleus like a swarm of angry bees,” as Singer terms it — until they happen to hit a pore in the nuclear membrane. Only then does the cell’s machinery lift a finger and actively shuttle mRNA through this gate.
More recently, Singer and colleagues created mutant mice that enabled them to watch as mRNA shuttled up and down a nerve cell’s delicate dendrites, the structures that receive signals from other nerves. The team even got to spy on memory-making in action. The mRNAs they were tracking carried instructions for making a protein — β-actin — that is abundant in nerve cells and is thought to help bolster connections when memories are made in the brain. In a video that looks like a network of roads at nighttime, within 10 minutes after a nerve cell was activated, mRNAs cruised to points of contact with other nerves, ready for actin production to shore up those nerve-nerve connections.
Scads of details about gene activity remain mysterious still, but it’s already clear that the process is far more dynamic than once assumed. “The change has been phenomenal, and it’s accelerating rapidly,” Singer says. “There’s a lot of information to be gleaned just by watching.”
This article originally appeared in Knowable Magazine, a nonprofit publication dedicated to making scientific knowledge accessible to all. Sign up for Knowable Magazine’s newsletter.